Published: Monday, May 20, 2020
Editor’s Notes
This article has been reviewed by Science X.
Editorial process
Policies.
Editors have highlighted
The content must have the following attributes to ensure its credibility:
fact-checked
Trusted source
Proofread
Chinese Academy of Sciences
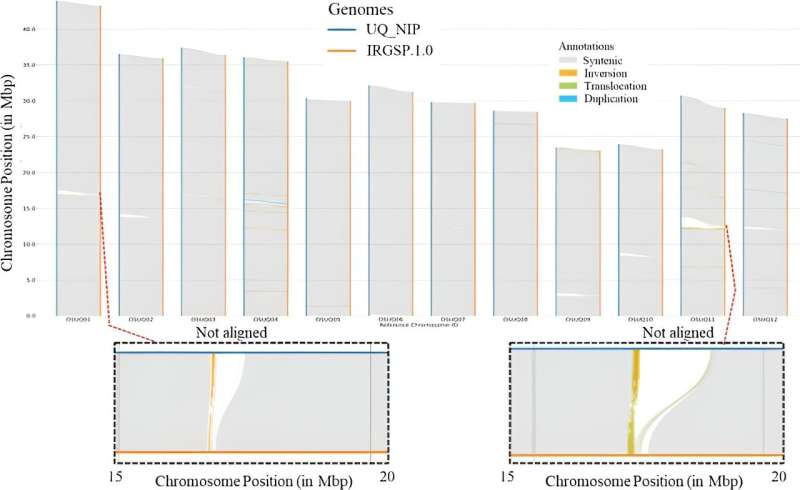
The IRGSP-1.0 Nipponbare assembly and UQ Nipponbare assembly were compared for sequence collinearity, structural variants (including inversions and translocations), and non-aligning areas. Tropical Plants 2024. DOI: 10.48130/tp-0024-0007
The haplotype resolved genome sequence for the japonica rice cultivar Nipponbare has been improved by a research team. This enhancement reveals more than 3,000 genes that have been identified and annotated. It could lead to significant improvements in crop improvement strategies and breeding.
Since its first sequencing, more than 20 years ago, the japonica rice Nipponbare cultivar has played a pivotal role in rice genomics. This was a major breakthrough in plant genomes. The Nipponbare genome assembly contains gaps despite continuous improvements in sequencing technologies, mostly due to repetitive DNA sequence.
Ongoing efforts have improved genome assembly for other rice species, and the technology has been extended to telomere sequence. The rice genome research has not yet addressed the problem of achieving a haplotype-resolved full assembly. This is a crucial area for future studies.
A study published on Tropical Plants, 3 April 2024 generates a rice genome with improved haplotype resolution for a comprehensive T2T improvement.
This study used PacBio HiFi and Hi-C to create a contig with Hifiasm. The result was a haplotype staged assembly. The assembly produced distinct contigs on nine of the chromosomes. The assembly of the remaining three chromosomes, however, involved the use of two contigs per chromosome.
The YaHS scaffolding software was used to organize the assembled contigs into 12 pseudochromosomes. This resulted in a T2T reference that was more complete and larger than the IRGSP-1.0. This refined assembly revealed 3,050 genes with transcript evidence supporting more than 95% of them, which indicates a significant improvement in genome annotation and structural understanding.
The findings demonstrate the enormous potential for new sequencing technologies to refine and expand genomic data. This will improve the genetic information in established genomes. The more detailed and extended genome that covers 99.3% universal single-copy gene with a N50 of 30.07 Mb provides a solid framework for future rice genetic studies and breeding programmes.
The comparative analyses also revealed structural variants as well as additional non-aligning areas, enriching our understanding of genomic architecture.
Robert J. Henry is the lead researcher of this study. He says that “this phased genome” will be an important resource for rice researchers. This team’s future plans include applying its advanced sequencing and assembling techniques to other varieties of rice and closely related species.
This work highlights the rapid development of rice genomics and the need for continued advancements in order to accurately map complex genomics.
For more information, please visit:
Muhammad Abdullah, et. al. An improved haplotype-resolved genome reveals additional rice genes. Tropical Plants, 2024. DOI: 10.48130/tp-0024-0007
Citation:
The rice genome research offers insights and applications for agriculture (2024 May 20).
Retrieved 20 May 2024
from https://phys.org/news/2024-05-advances-rice-genome-insights-applications.html
Copyright applies to this document. This document is protected by copyright.
No part of the website may be reproduced or copied without written permission. This content is only provided as information.
Source: Phys.org



































